Starting with Data
Last updated on 2024-03-12 | Edit this page
Estimated time: 80 minutes
Overview
Questions
- What is a data.frame?
- How can I read a complete csv file into R?
- How can I get basic summary information about my dataset?
- How can I change the way R treats strings in my dataset?
- Why would I want strings to be treated differently?
- How are dates represented in R and how can I change the format?
Objectives
- Describe what a data frame is.
- Load external data from a .csv file into a data frame.
- Summarize the contents of a data frame.
- Describe the difference between a factor and a string.
- Convert between strings and factors.
- Reorder and rename factors.
- Change how character strings are handled in a data frame.
- Examine and change date formats.
Open your Rproj file
First, open your R Project file
(library_carpentry.Rproj) created in the
Before We Start lesson.
If you did not complete that step, do the following:
- Under the
Filemenu, click onNew project, chooseNew directory, thenNew project - Enter the name
library_carpentryfor this new folder (or “directory”). This will be your working directory for the rest of the day. - Click on
Create project - Create a new file where we will type our scripts. Go to File >
New File > R script. Click the save icon on your toolbar and save
your script as “
script.R”.
Presentation of the data
This data was downloaded from the University of Houston–Clear Lake Integrated Library System in 2018. It is a relatively random sample of books from the catalog. It consists of 10,000 observations of 11 variables.
These variables are:
-
CALL...BIBLIO.: Bibliographic call number. Most of these are cataloged with the Library of Congress classification, but there are also items cataloged in the Dewey Decimal System (including fiction and non-fiction), and Superintendent of Documents call numbers. Character. -
X245.ab: The title and remainder of title. Exported from MARC tag 245|ab fields. Separated by a|pipe character. Character. -
X245.c: The author (statement of responsibility). Exported from MARC tag 245|c. Character. -
TOT.CHKOUT: The total number of checkouts. Integer. -
LOUTDATE: The last date the item was checked out. Date. YYYY-MM-DDThh:mmTZD -
SUBJECT: Bibliographic subject in Library of Congress Subject Headings. Separated by a|pipe character. Character. -
ISN: ISBN or ISSN. Exported from MARC field 020|a. Character -
CALL...ITEM: Item call number. Most of these areNAbut there are some secondary call numbers. -
X008.Date.One: Date of publication. Date. YYYY -
BCODE2: Item format. Character. -
BCODE1Sub-collection. Character.
Getting data into R
Ways to get data into R
In order to use your data in R, you must import it and turn it into an R object. There are many ways to get data into R.
-
Manually: You can manually create it using the
data.frame()function in Base R, or thetibble()function in the tidyverse. -
Import it from a file Below is a very incomplete
list
- Text: TXT (
readLines()function) - Tabular data: CSV, TSV (
read.table()function orreadrpackage) - Excel: XLSX (
xlsxpackage) - Google sheets: (
googlesheetspackage) - Statistics program: SPSS, SAS (
havenpackage) - Databases: MySQL (
RMySQLpackage)
- Text: TXT (
-
Gather it from the web: You can connect to
webpages, servers, or APIs directly from within R, or you can create a
data scraped from HTML webpages using the
rvestpackage. For example
Organizing your working directory
Using a consistent folder structure across your projects will help keep things organized and make it easy to find/file things in the future. This can be especially helpful when you have multiple projects. In general, you might create directories (folders) for scripts, data, and documents. Here are some examples of suggested directories:
-
data/Use this folder to store your raw data and intermediate datasets. For the sake of transparency and provenance, you should always keep a copy of your raw data accessible and do as much of your data cleanup and preprocessing programmatically (i.e., with scripts, rather than manually) as possible. -
data_output/When you need to modify your raw data, it might be useful to store the modified versions of the datasets in a different folder. -
documents/Used for outlines, drafts, and other text. -
fig_output/This folder can store the graphics that are generated by your scripts. -
scripts/A place to keep your R scripts for different analyses or plotting.
You may want additional directories or subdirectories depending on your project needs, but these should form the backbone of your working directory.
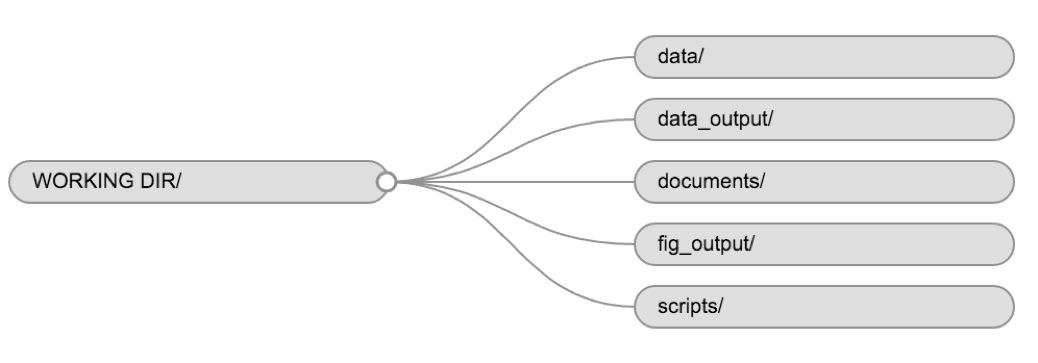
The working directory
The working directory is an important concept to understand. It is the place on your computer where R will look for and save files. When you write code for your project, your scripts should refer to files in relation to the root of your working directory and only to files within this structure.
Using RStudio projects makes this easy and ensures that your working
directory is set up properly. If you need to check it, you can use
getwd(). If for some reason your working directory is not
what it should be, you can change it in the RStudio interface by
navigating in the file browser to where your working directory should
be, then by clicking on the blue gear icon “More”, and selecting “Set As
Working Directory”. Alternatively, you can use
setwd("/path/to/working/directory") to reset your working
directory. However, your scripts should not include this line, because
it will fail on someone else’s computer.
Setting your working directory withsetwd()
Some points to note about setting your working directory:
The directory must be in quotation marks.
On Windows computers, directories in file paths are separated with a
backslash \. However, in R, you must use a forward slash
/. You can copy and paste from the Windows Explorer window
directly into R and use find/replace (Ctrl/Cmd + F) in R Studio to
replace all backslashes with forward slashes.
On Mac computers, open the Finder and navigate to the directory you
wish to set as your working directory. Right click on that folder and
press the options key on your keyboard. The ‘Copy “Folder
Name”’ option will transform into ’Copy “Folder Name” as Pathname. It
will copy the path to the folder to the clipboard. You can then paste
this into your setwd() function. You do not need to replace
backslashes with forward slashes.
After you set your working directory, you can use ./ to
represent it. So if you have a folder in your directory called
data, you can use read.csv(“./data”) to represent that
sub-directory.
Downloading the data and getting set up
Now that you have set your working directory, we will create our
folder structure using the dir.create() function.
For this lesson we will use the following folders in our working
directory: data/,
data_output/ and
fig_output/. Let’s write them all in
lowercase to be consistent. We can create them using the RStudio
interface by clicking on the “New Folder” button in the file pane
(bottom right), or directly from R by typing at console:
R
dir.create("data")
dir.create("data_output")
dir.create("fig_output")
Go to the Figshare page for this curriculum and download the dataset
called “books.csv”. The direct download link is: https://ndownloader.figshare.com/files/22031487.
Place this downloaded file in the data/ you just created.
Alternatively, you can do this directly from R by copying and
pasting this in your terminal (your instructor can place this chunk of
code in the Etherpad):
R
download.file("https://ndownloader.figshare.com/files/22031487",
"data/books.csv", mode = "wb")
Now if you navigate to your data folder, the
books.csv file should be there. We now need to load it into
our R session.
tidyverse
R has some base functions for reading a local data file into your R
session–namely read.table() and read.csv(),
but these have some idiosyncrasies that were improved upon in the
readr package, which is installed and loaded with
tidyverse.
R
library(tidyverse) # loads the core tidyverse, including dplyr, readr, ggplot2, purrr
OUTPUT
── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ dplyr 1.1.2 ✔ purrr 1.0.1
✔ forcats 1.0.0 ✔ stringr 1.5.0
✔ ggplot2 3.4.2 ✔ tibble 3.2.1
✔ lubridate 1.9.2 ✔ tidyr 1.3.0
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorsTo get our sample data into our R session, we will use the
read_csv() function and assign it to the books
value.
R
books <- read_csv("./data/books.csv")
You will see the message
Parsed with column specification, followed by each column
name and its data type. When you execute read_csv on a data
file, it looks through the first 1000 rows of each column and guesses
the data type for each column as it reads it into R. For example, in
this dataset, it reads SUBJECT as
col_character (character), and TOT.CHKOUT as
col_double. You have the option to specify the data type
for a column manually by using the col_types argument in
read_csv.
You should now have an R object called books in the
Environment pane: 10000 observations of 12 variables. We will be using
this data file in the next module.
Note
read_csv() assumes that fields are delineated by commas,
however, in several countries, the comma is used as a decimal separator
and the semicolon (;) is used as a field delineator. If you want to read
in this type of files in R, you can use the read_csv2
function. It behaves exactly like read_csv but uses
different parameters for the decimal and the field separators. If you
are working with another format, they can be both specified by the user.
Check out the help for read_csv() by typing
?read_csv to learn more. There is also the
read_tsv() for tab-separated data files, and
read_delim() allows you to specify more details about the
structure of your file.
What are data frames and tibbles?
Data frames are the de facto data structure for tabular data
in R, and what we use for data processing, statistics, and
plotting.
A data frame is the representation of data in the format of a table where the columns are vectors that all have the same length. Because columns are vectors, each column must contain a single type of data (e.g., characters, integers, factors). For example, here is a figure depicting a data frame comprising a numeric, a character, and a logical vector.
A data frame can be created by hand, but most commonly they are
generated by the functions read_csv() or
read_table(); in other words, when importing spreadsheets
from your hard drive (or the web).
A tibble is an extension of R data frames used by the
tidyverse. When the data is read using read_csv(),
it is stored in an object of class tbl_df,
tbl, and data.frame. You can see the class of
an object with class().
Inspecting data frames
When calling a tbl_df object (like books
here), there is already a lot of information about our data frame being
displayed such as the number of rows, the number of columns, the names
of the columns, and as we just saw the class of data stored in each
column. However, there are functions to extract this information from
data frames. Here is a non-exhaustive list of some of these functions.
Let’s try them out!
-
Size:
-
dim(books)- returns a vector with the number of rows in the first element, and the number of columns as the second element (the dimensions of the object) -
nrow(books)- returns the number of rows -
ncol(books)- returns the number of columns
-
-
Content:
-
head(books)- shows the first 6 rows -
tail(books)- shows the last 6 rows
-
-
Names:
-
names(books)- returns the column names (synonym ofcolnames()fordata.frameobjects)
-
-
Summary:
-
View(books)- look at the data in the viewer -
str(books)- structure of the object and information about the class, length and content of each column -
summary(books)- summary statistics for each column
-
Note: most of these functions are “generic”, they can be used on other types of objects besides data frames.
The map() function from purrr is a useful
way of running a function on all variables in a data frame or list. If
you loaded the tidyverse at the beginning of the session,
you also loaded purrr. Here we call class() on
books using map_chr(), which will return a
character vector of the classes for each variable.
R
map_chr(books, class)
OUTPUT
CALL...BIBLIO. X245.ab X245.c LOCATION TOT.CHKOUT
"character" "character" "character" "character" "numeric"
LOUTDATE SUBJECT ISN CALL...ITEM. X008.Date.One
"character" "character" "character" "character" "character"
BCODE2 BCODE1
"character" "character" Indexing and subsetting data frames
Our books data frame has 2 dimensions: rows
(observations) and columns (variables). If we want to extract some
specific data from it, we need to specify the “coordinates” we want from
it. In the last session, we used square brackets [ ] to
subset values from vectors. Here we will do the same thing for data
frames, but we can now add a second dimension. Row numbers come first,
followed by column numbers. However, note that different ways of
specifying these coordinates lead to results with different classes.
R
## first element in the first column of the data frame (as a vector)
books[1, 1]
## first element in the 6th column (as a vector)
books[1, 6]
## first column of the data frame (as a vector)
books[[1]]
## first column of the data frame (as a data.frame)
books[1]
## first three elements in the 7th column (as a vector)
books[1:3, 7]
## the 3rd row of the data frame (as a data.frame)
books[3, ]
## equivalent to head_books <- head(books)
head_books <- books[1:6, ]
Dollar sign
The dollar sign $ is used to distinguish a specific
variable (column, in Excel-speak) in a data frame:
R
head(books$X245.ab) # print the first six book titles
OUTPUT
[1] "Bermuda Triangle /"
[2] "Invaders from outer space :|real-life stories of UFOs /"
[3] "Down Cut Shin Creek :|the pack horse librarians of Kentucky /"
[4] "The Chinese book of animal powers /"
[5] "Judge Judy Sheindlin's Win or lose by how you choose! /"
[6] "Judge Judy Sheindlin's You can't judge a book by its cover :|cool rules for school /"R
# print the mean number of checkouts
mean(books$TOT.CHKOUT)
OUTPUT
[1] 2.2847
unique(), table(), and
duplicated()
Use unique() to see all the distinct values in a
variable:
R
unique(books$BCODE2)
OUTPUT
[1] "a" "w" "s" "m" "e" "4" "k" "5" "n" "o"Take that one step further with table() to get quick
frequency counts on a variable:
R
table(books$BCODE2) # frequency counts on a variable
OUTPUT
4 5 a e k m n o s w
1 3 6983 68 3 109 2 21 1988 822 You can combine table() with relational operators:
R
table(books$TOT.CHKOUT > 50) # how many books have 50 or more checkouts?
OUTPUT
FALSE TRUE
9991 9 duplicated() will give you the a logical vector of
duplicated values.
R
duplicated(books$ISN) # a TRUE/FALSE vector of duplicated values in the ISN column
!duplicated(books$ISN) # you can put an exclamation mark before it to get non-duplicated values
table(duplicated(books$ISN)) # run a table of duplicated values
which(duplicated(books$ISN)) # get row numbers of duplicated values
Exploring missing values
You may also need to know the number of missing values:
R
sum(is.na(books)) # How many total missing values?
OUTPUT
[1] 14509R
colSums(is.na(books)) # Total missing values per column
OUTPUT
CALL...BIBLIO. X245.ab X245.c LOCATION TOT.CHKOUT
561 12 2801 0 0
LOUTDATE SUBJECT ISN CALL...ITEM. X008.Date.One
0 63 2934 7980 158
BCODE2 BCODE1
0 0 R
table(is.na(books$ISN)) # use table() and is.na() in combination
OUTPUT
FALSE TRUE
7066 2934 R
booksNoNA <- na.omit(books) # Return only observations that have no missing values
Exercise
Call
View(books)to examine the data frame. Use the small arrow buttons in the variable name to sort tot_chkout by the highest checkouts. What item has the most checkouts?What is the class of the TOT.CHKOUT variable?
Use
table()andis.na()to find out how many NA values are in the ISN variable.Call
summary(books$TOT.CHKOUT). What can we infer when we compare the mean, median, and max?hist()will print a rudimentary histogram, which displays frequency counts. Callhist(books$TOT.CHKOUT). What is this telling us?
Highest checkouts:
Click, clack, moo : cows that type.class(books$TOT.CHKOUT)returnsnumerictable(is.na(books$ISN))returns 2934TRUEvaluesThe median is 0, indicating that, consistent with all book circulation I have seen, the majority of items have 0 checkouts.
As we saw in
summary(), the majority of items have a small number of checkouts
Logical tests
R contains a number of operators you can use to compare values. Use
help(Comparison) to read the R help file. Note that
two equal signs (==) are used for
evaluating equality (because one equals sign (=) is used
for assigning variables).
| operator | function |
|---|---|
< |
Less Than |
> |
Greater Than |
== |
Equal To |
<= |
Less Than or Equal To |
>= |
Greater Than or Equal To |
!= |
Not Equal To |
%in% |
Has a Match In |
is.na() |
Is NA |
!is.na() |
Is Not NA |
Sometimes you need to do multiple logical tests (think Boolean
logic). Use help(Logic) to read the help file.
| operator | function |
|---|---|
& |
boolean AND |
| ` | ` |
! |
Boolean NOT |
any() |
Are some values true? |
all() |
Are all values true? |
